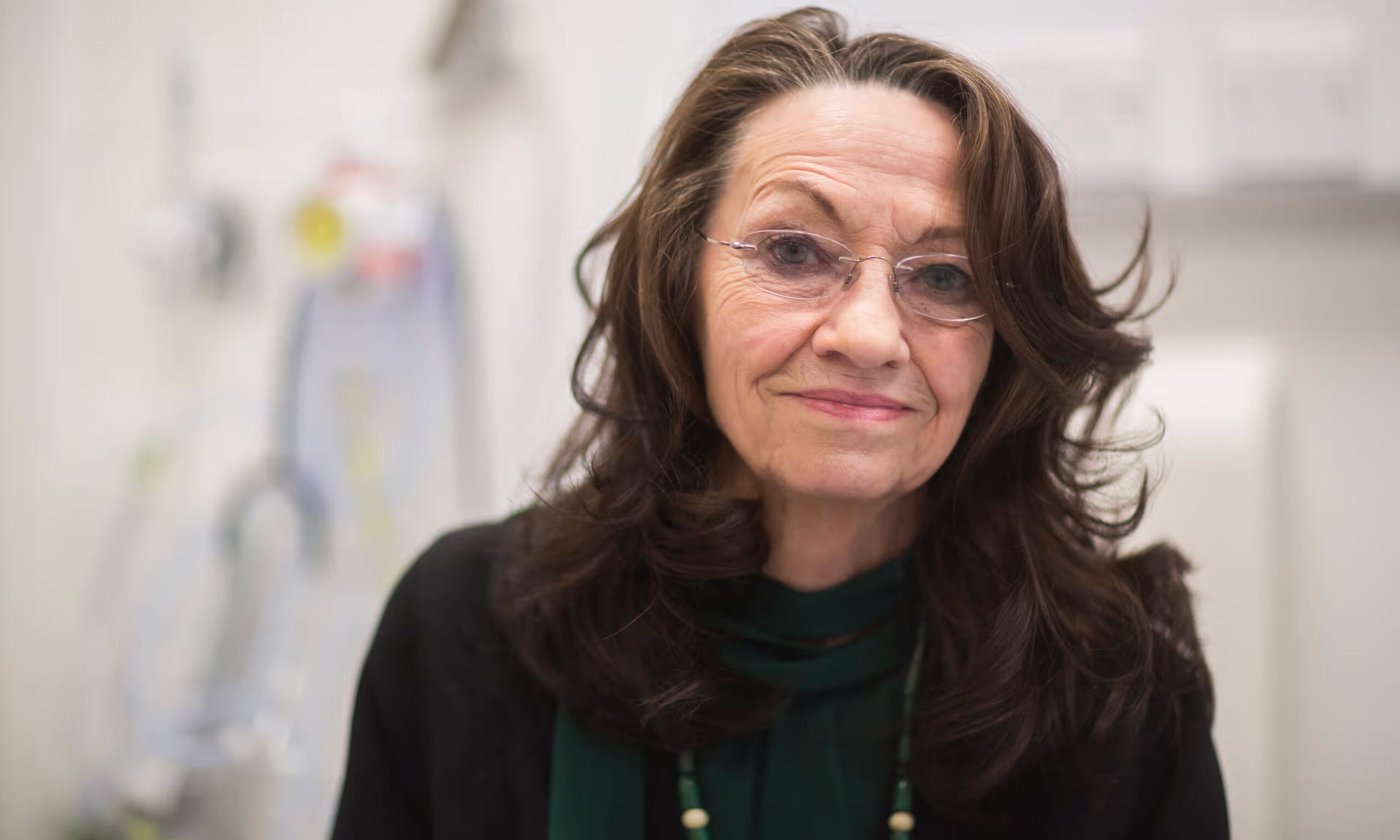
Rosenfeld Group
Liquid biopsies and cancer diagnostics
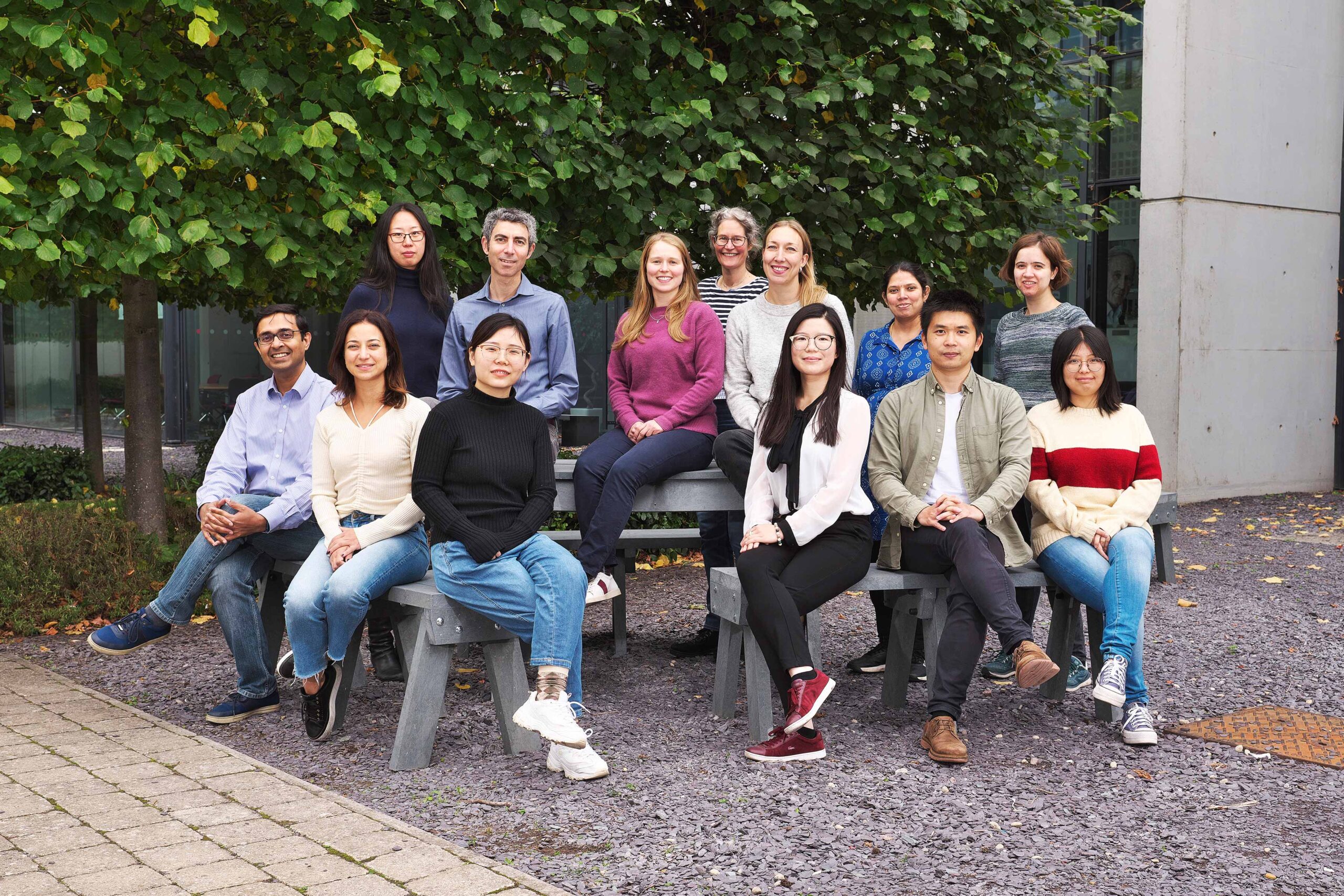

Research summary
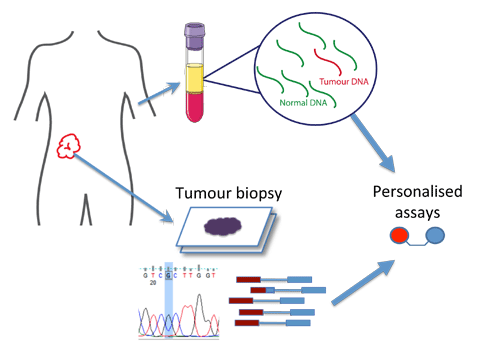
Cancer patients’ blood contains DNA fragments from tumors. Using advanced genomic technologies, we have developed liquid biopsies to help diagnose cancer and allow us to study its evolution and treatment resistance noninvasively. This method could offer more detailed and frequent monitoring compared to traditional tissue biopsies, leading to better research and improved cancer care.

Professor Nitzan Rosenfeld
Senior Group Leader,
Director of Barts Cancer Institute
Research areas
Related News
See all news-
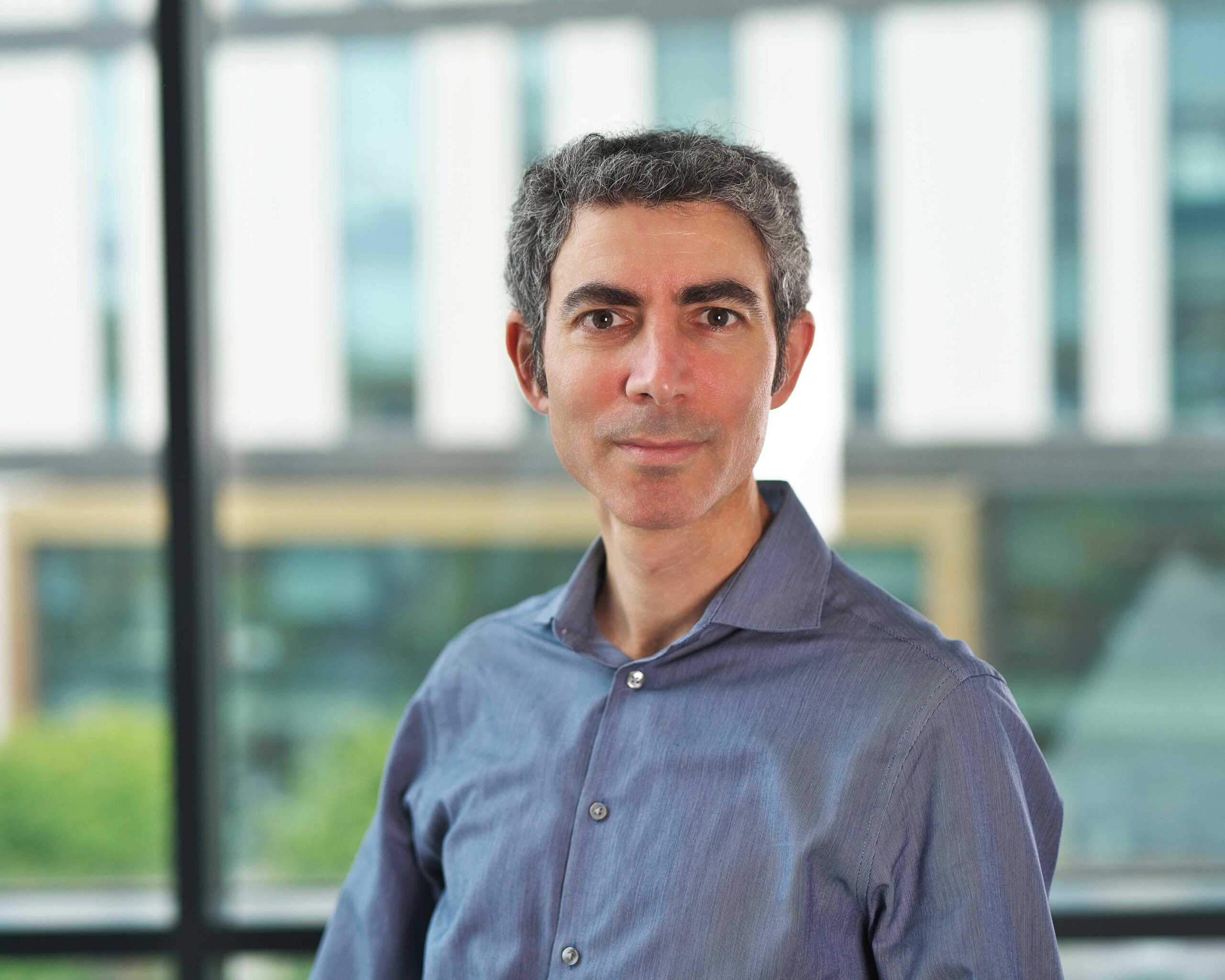

Prof Nitzan Rosenfeld appointed Director of Barts Cancer Institute
22nd September 2023
Professor Nitzan Rosenfeld has been appointed as Queen Mary University of London’s new Director of the Barts Cancer Institute.
Find out more -

Institute Group Leaders awarded promotions by the University
15th June 2022
We are delighted to announce that Florian Markowetz and Nitzan Rosenfeld have been promoted to Professors in the University of Cambridge Senior Academic Promotions.
Find out more -
Personalised blood test can detect persistent lung cancer
17th March 2022
Patients who are at a higher risk of their lung cancer returning can be identified by a personalised blood test that is performed after treatment, according to researchers at the University of Cambridge.
Find out more
Laboratory Efficiency Assessment Framework (LEAF)
The Rosenfeld Group contributed to the Institute’s LEAF Silver accreditation, see the Sustainability webpage for more information.