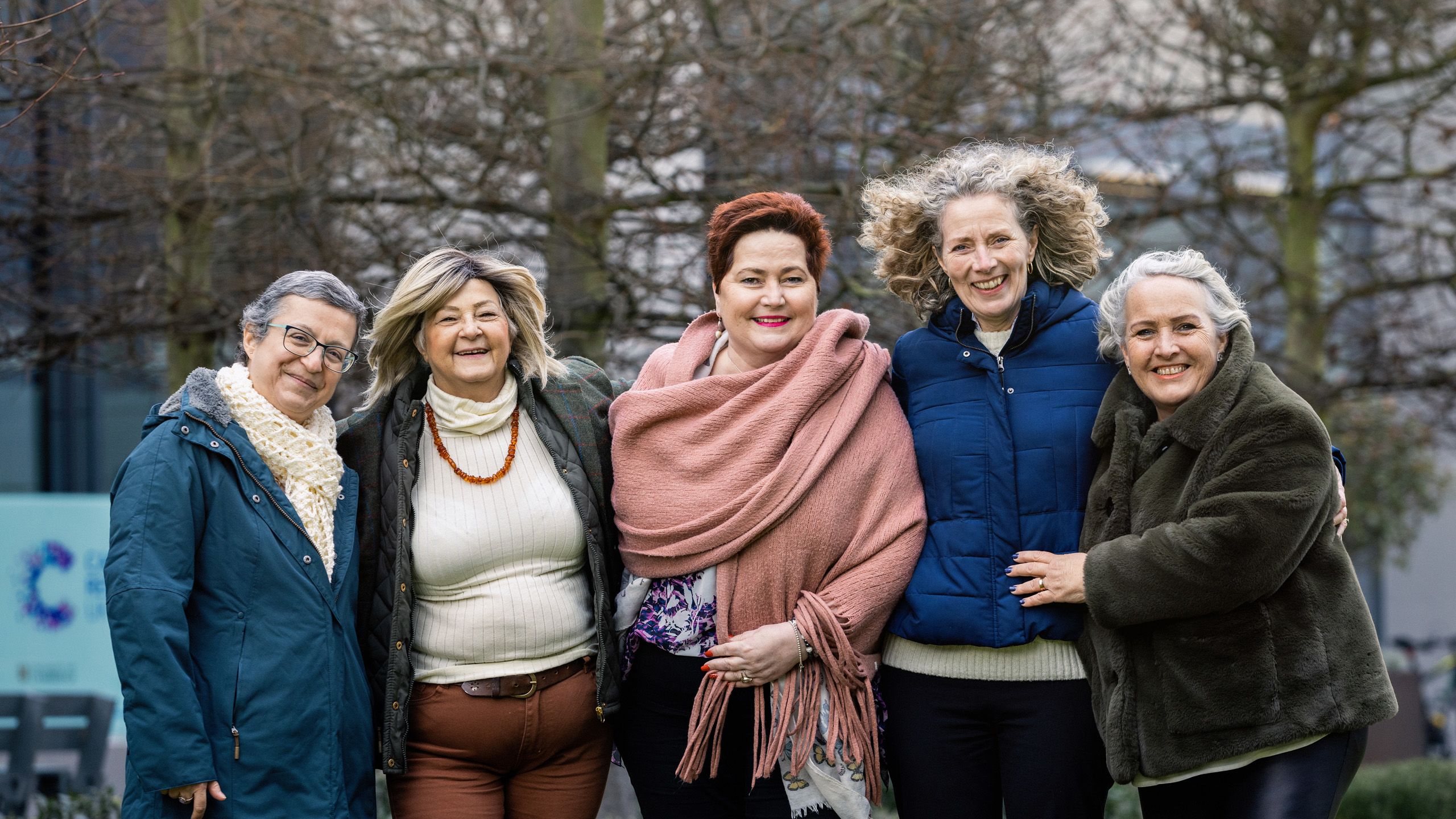
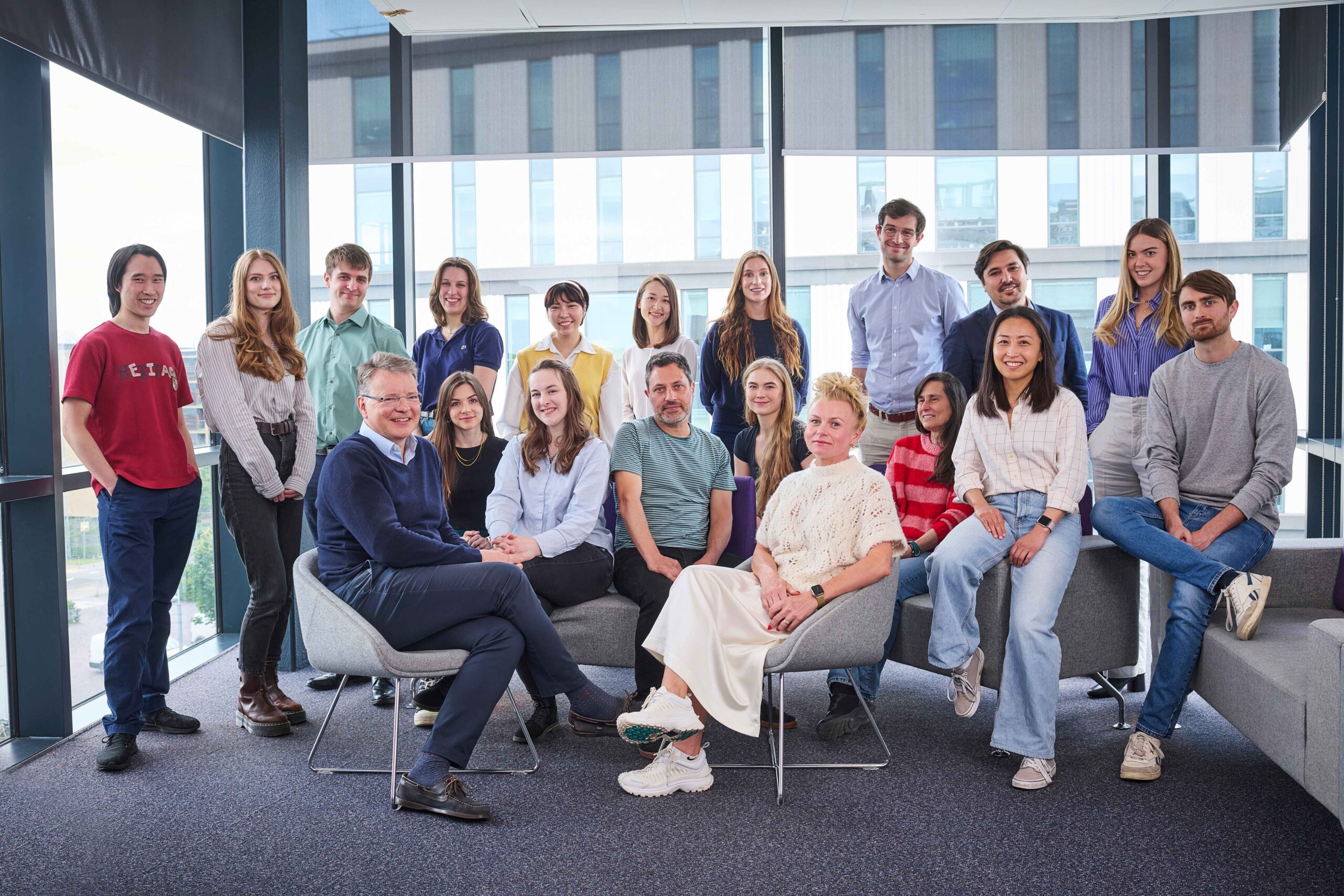
Brenton Group
Functional genomics of ovarian cancer

Research summary
We study the genetic changes that happen in ovarian cancer to understand what causes cancer cells to become resistant to treatment. We use medical imaging technology to study how tumours respond to treatment and also to help us study cancers that have spread to other parts of the body. By understanding how drug resistance develops, we hope to improve treatments by tailoring drug regimes according to the genetic ‘signature’ of a patient’s cancer.
Introduction
Our laboratory focuses on discovering improved treatments for epithelial ovarian cancer using laboratory and clinical studies.

Professor James Brenton
Senior Group Leader
Research topics
Related News
See all news-
1M to advance AI powered personalised ovarian cancer care
19th February 2026
Researchers from our Brenton Group are part of an international team awarded the Global Ovarian Cancer Research Consortium’s inaugural AI Accelerator Grant.
Find out more -
Targeting paused cells could improve chemotherapy for lung and ovarian cancers
3rd February 2026
New research published today in Nature Aging by scientists at the University of Cambridge sheds light on why some lung and ovarian cancers stop responding to chemotherapy, and how this resistance might one day be prevented.
Find out more -
Single-cell study sheds new light on why ovarian cancer becomes resistant to chemotherapy
11th August 2025
Researchers at the Cancer Research UK Cambridge Institute and Stanford University have mapped how ovarian cancer cells respond to chemotherapy at an unprecedented level of detail, offering new insights into why treatment resistance develops.
Find out more
Publications
See All Publications-
Rare germline genetic variation in PAX8 transcription factor binding sites and susceptibility to epithelial ovarian cancer.
Brenton Group
E-pub date: 8 Apr 2025
-
PTEN Loss Shapes Macrophage Dynamics in High-Grade Serous Ovarian Carcinoma.
Brenton Group
E-pub date: 15 Nov 2024
Laboratory Efficiency Assessment Framework (LEAF)
The Brenton Group contributed to the Institute’s LEAF Silver accreditation, see the Sustainability webpage for more information.